Could computers of the future run on human brain cells? A team of researchers at Johns Hopkins University certainly think so. In a paper published in the journal Frontiers in Science, the team outline their plans for ‘organoid intelligence’, an emerging multidisciplinary field looking to develop biocomputers that operate with human brains cells. Such a development could not only massively expand the capabilities of modern computing but also open up new fields of study.
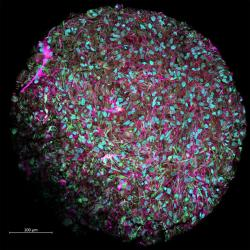
Organoids are tiny, self-organizing 3D tissues that are typically derived from stem cells, and mimic the main functional and architectural complexity of an organ. It is possible there could be as many types of organoids as there are tissues and organs in the body. To date, scientists have produced organoid cultures for intestines, liver, pancreas, kidneys, prostate, lung, optic cup, and the brain, and it seems more may be on the way.
These tissues provide unique opportunities for scientists to study human diseases that do not rely on traditional methods associated with animal models. The reliance on animal models has historically led to a bottleneck in treatment discovery as there are biological processes that are specific to the human body and cannot be modeled on animals. The development of organoids promises to overcome these limitations. Yet the team at Johns Hopkins University are taking the research into organoids in a completely different direction.
“Computing and artificial intelligence have been driving the technology revolution but they are reaching a ceiling,” explained Thomas Hartung, a professor of environmental health sciences at the Johns Hopkins Bloomberg School of Public Health and Whiting School of Engineering, in a statement. “Biocomputing is an enormous effort of compacting computational power and increasing its efficiency to push past our current technological limits.”
Sources:
https://www.frontiersin.org/journals/science/articles/10.3389/fsci.2023.1017235
https://www.iflscience.com/is-oi-the-new-ai-biocomputers-could-one-day-run-on-human-brain-cells-67732
https://www.osel.cz/12752-zapomente-na-elektronicke-chatboty-rysuji-se-organoidni-inteligence.html